

These software packages are written and maintained by the lab. More information on request.
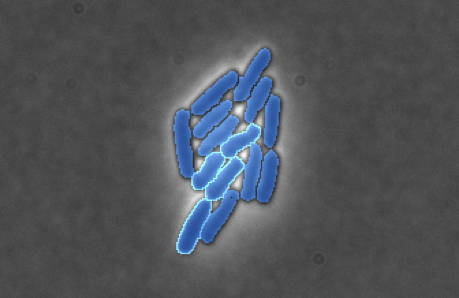
SuperSegger & SuperSegger-Omniposeis a completely automated MATLAB-based trainable image cell segmentation, fluorescence quantification and analysis suite, particularly well suited for high-throughput time lapse fluorescence microscopy of in vivo bacterial cells.NEW! SuperSegger has also been modified to work with the improved segmentation of Omnipose. Supersegger-Omnipose can be found here. The original SuperSegger website can be found here, with the Github here.
|  |

|
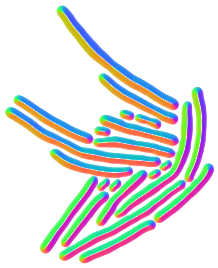
Omniposesolves the over-segmentation problems of Cellpose, a generalist algorithm for cell and nucleus segmentation, for large, anisotropic cells. This is particularly relevant for bacterial cells, but Omnipose is suitable for arbitrary cell shapes. Click here to download. |
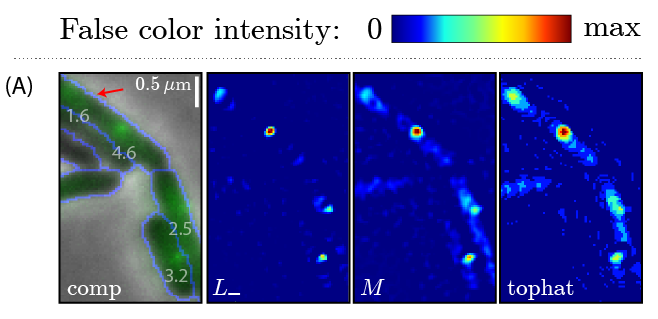
Curve Filteris a matlab-based tool for finding punctate foci in fluorescent microscopy images.Click here to download. |
 |
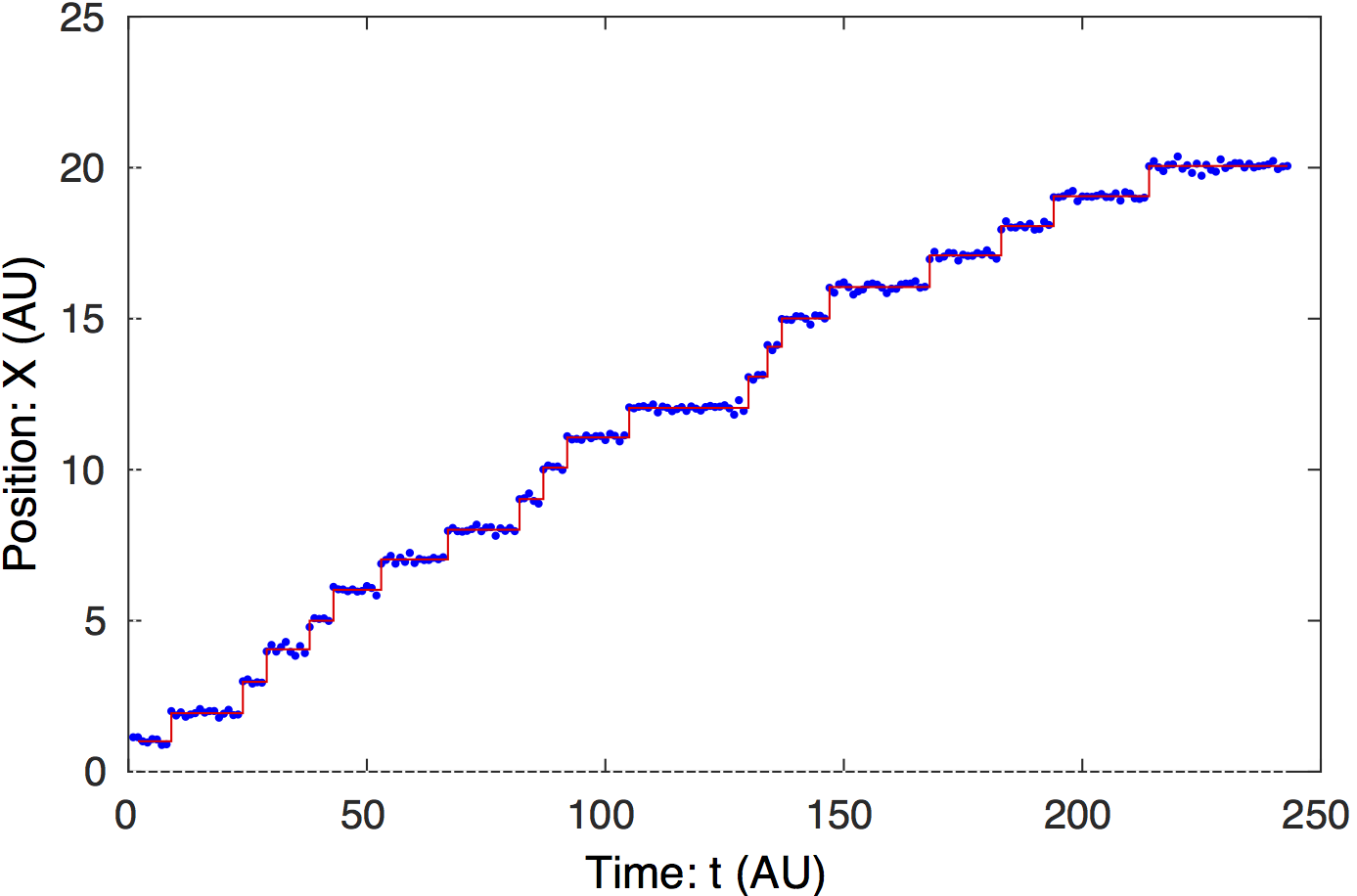 |
Steppiis a matlab-based tool for change-point analysis. We present a unified approach to the analysis of processes whose noise can be modeled by Gaussian, Wiener or Ornstein-Uhlenbeck Processes. We expect this technique to be of general interest to experimental investigators interested in biological systems. Applications of this analysis include molecular-motor stepping, fluorophore bleaching, electrophysiology, particle and cell tracking, detection of copy number variation by sequencing, tethered- particle-motion etc.Click here for information and download. |
Wormulatoris a DNA statistics calculator that can be used with a web-based interface. Click here to view it. | 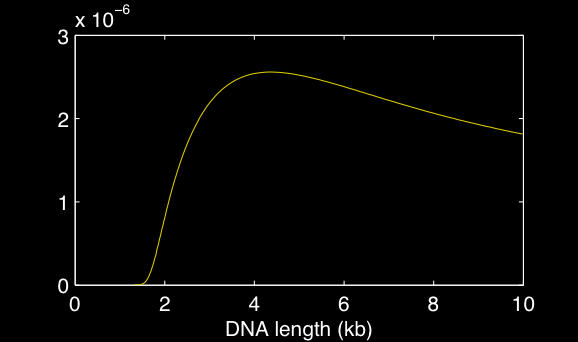 |
 |
Traceis a matlab-based polymer tracking software package which we have used for tracing DNA contours. Click here to download. |