| Home | Gene Index | Wiggins Lab | Invert Page |
uracil phosphoribosyltransferase
Gene Ontology: | |
| GO:0000166 | nucleotide binding |
| GO:0003824 | catalytic activity |
| GO:0004845 | uracil phosphoribosyltransferase activity |
| GO:0005525 | GTP binding |
| GO:0005829 | cytosol |
| GO:0006223 | uracil salvage |
| GO:0008152 | metabolic process |
| GO:0009116 | nucleoside metabolic process |
| GO:0016020 | membrane |
| GO:0016740 | transferase activity |
| GO:0016757 | transferase activity, transferring glycosyl groups |
| GO:0042802 | identical protein binding |
| GO:0044206 | UMP salvage |
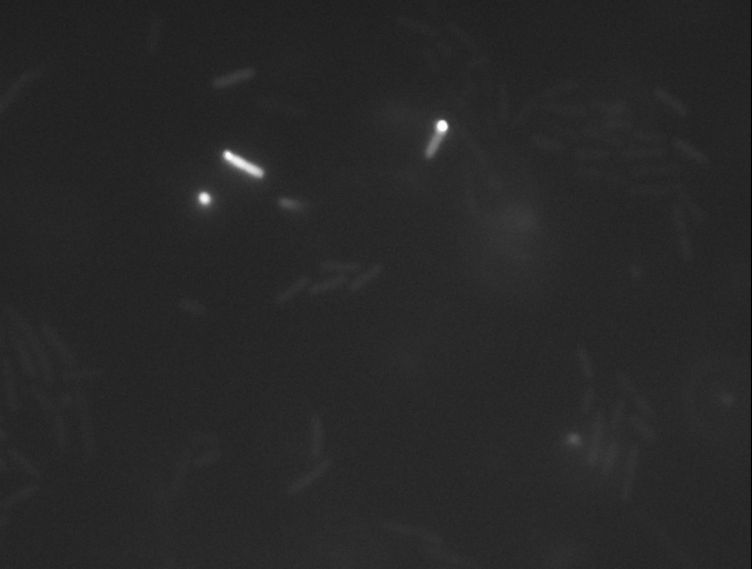
Protein localization Map: upp | |||||||
| Gene name: | upp | Gene name: | upp | Gene name: | upp | ||
| Strain: | JW2483 | Strain: | JW2483 | Strain: | JW2483 | ||
| Induction: | 500 uM IPTG | Induction: | 500 uM IPTG | Induction: | 50 uM IPTG | ||
| Comment: | 7A2F | Comment: | 7A22F | Comment: | 7A2F | ||
| Reference: | PMID:16769691--ASKA Collection--Kitagawa et al. | Reference: | PMID:16769691--ASKA Collection--Kitagawa et al. | Reference: | PMID:16769691--ASKA Collection--Kitagawa et al. | ||
|
|
|
Cell Orientation
|
Relative Intensity
|
|||
Most similar localization patterns. gene: distance | ||||
|
|
|
|
|
|
|
|
|
|