| Home | Gene Index | Wiggins Lab | Invert Page |
"exonuclease, dsDNA, ATP-dependent"
Gene Ontology: | |
| GO:0000166 | nucleotide binding |
| GO:0000725 | recombinational repair |
| GO:0004519 | endonuclease activity |
| GO:0005524 | ATP binding |
| GO:0006260 | DNA replication |
| GO:0008296 | 3'-5'-exodeoxyribonuclease activity |
| GO:0090305 | nucleic acid phosphodiester bond hydrolysis |
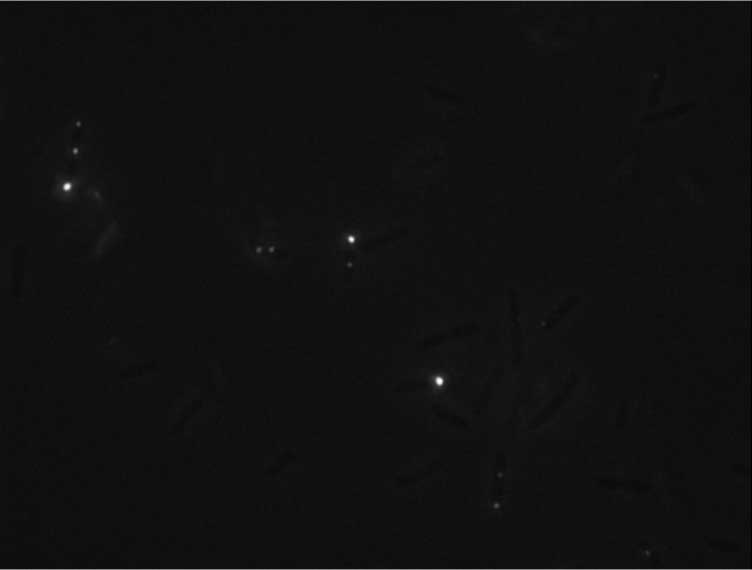
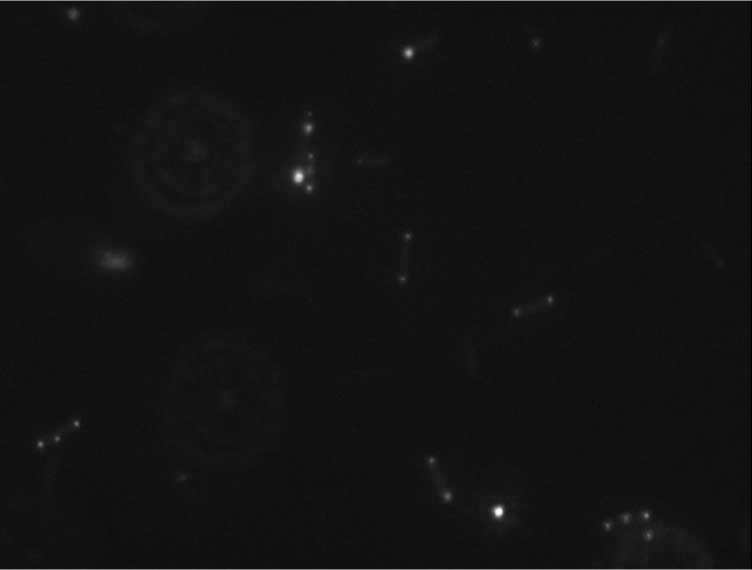
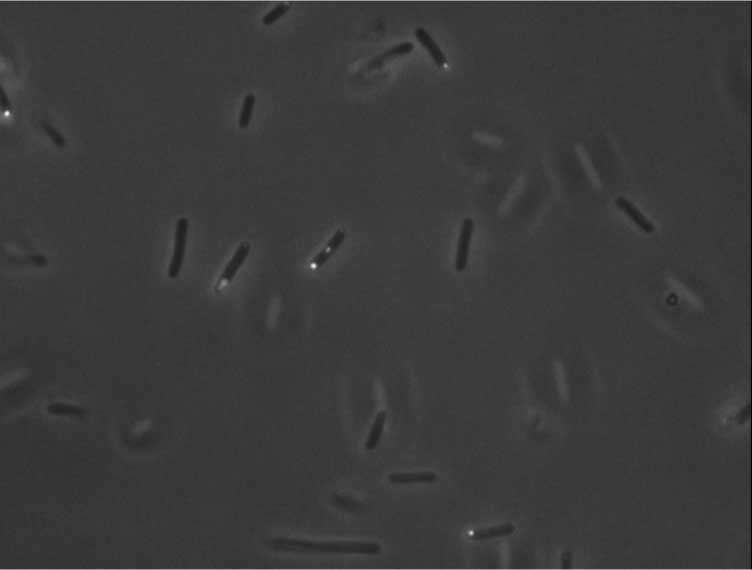
Protein localization Map: sbcC | |||
| Gene name: | sbcC | ||
| Strain: | JW0387 | ||
| Induction: | 50 uM IPTG | ||
| Comment: | 3B8E | ||
| Reference: | PMID:16769691--ASKA Collection--Kitagawa et al. | ||
|
Cell Orientation
|
Relative Intensity
|
|
Most similar localization patterns. gene: distance | ||||
|
|
|
|
|
|
|
|
|
|